DNA helix structure
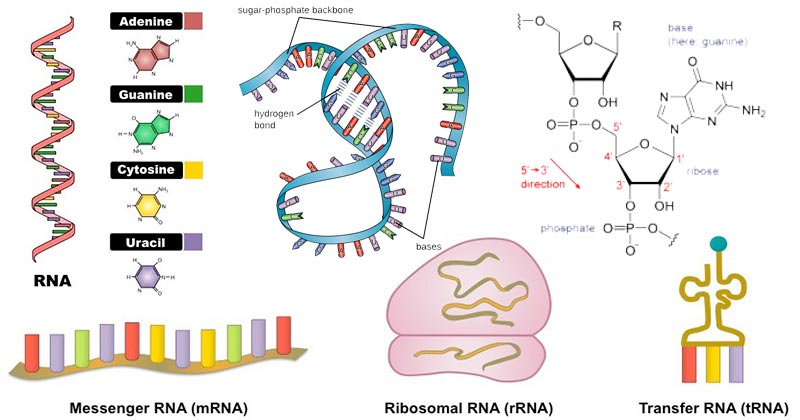
There are two types of nucleic acids namely: DNA- Deoxyribonucleic. RNA - Ribonucleic acid.
Localization
In prokaryotes both transcription and translation occur in the cytoplasm due to the absence of nucleus. In eukaryote transcription occurs in the nucleus and translation occurs in ribosomes present on the rough endoplasmic membrane in the cytoplasm.
Factors
Transcription is performed by RNA polymerase and other associated proteins termed as transcription factors. It can be inducible as seen in the spatio-temporal regulation of developmental genes or consitutive as seen in case of house keeping genes like Gapdh.
Translation is performed by a multisubunit structure called ribosome which consists of rRNA and proteins.
Initiation
Transcription initiates with RNA polymerase binding to the promoter region in the DNA. The transcription factors and RNA polymerase binding to the promoter forms a transcription initiation complex. The promoter consists of a core region like the TATA box where the complex binds. It is in this stage that RNA polymerase unwinds the DNA.
Translation initiates with the formation of initiation complex. The ribosome subunit, three initiation factors (IF1, IF2 and IF3) and methionine carrying t-RNA bind the mRNA near the AUG start codon.
Elongation
During transcription, the RNA polymerase after the initial abortive attempts traverses the template strand of DNA in 3’ to 5’ direction, producing a complementary RNA strand in 5’ to 3’ direction. As the RNA polymerase advances the DNA strand that has been transcribed rewinds to form a double helix.
During translation the incoming aminoacyl t-RNA binds to the codon (sequences of 3 nucleotides) at A-site and a peptide bond is formed between the new amino acid and the growing chain. The peptide then moves one codon position to get ready for the next amino acid. The process hence proceeds in a 5’ to 3’ direction.
Termination
Transcription termination in prokaryotes can either be Rho-independent, where a GC rich hairpin loop is formed or Rho-dependent, where a protein factor Rho destabilizes the DNA-RNA interaction. In eukaryotes when a termination sequence is encountered the RNA nascent transcript is released and it is poly-adenylated.
In translation when the ribosome encounters one of the three stop codons it disassembles the ribosome and releases the polypeptide.
End Product
The end product of transcription is an RNA transcript which can form any of the following types of RNA: mRNA, tRNA, rRNA and non-coding RNA (like microRNA). Usually in prokaryotes the mRNA formed is polycistronic and in eukaryotes it is monocistronic.
The end product of translation is a polypeptide chain which folds and undergoes post translational modifications to form a functional protein.
Post Process Modification
During post transcriptional modification in eukaryotes, a 5’ cap, a 3’ poly tail is added and introns are spliced out. In prokaryotes this process is absent.
A number of post-translational modifications occur including phosphorylation, SUMOylation, disulfide bridges formation, farnesylation etc.
Antibiotics
Transcription is inhibited by rifampicin (antibacterial) and 8-Hydroxyquinoline (antifungal).
Translation is inhibited by anisomycin, cycloheximide, chloramphenicol, tetracyclin, streptomycin, erythromycin and puromycin.
Methods to measure and detect
For Transcription, RT-PCR, DNA microarray, In-situ hybridization, Northern blot, RNA-Seq is quite often used for measurement and detection.For Translation, western blotting, immunoblotting, enzyme assay, Protein sequencing, Metabolic labeling, proteomics is used for measurement and detection.
Crick’s central dogma: DNA ---> Transcription ---> RNA ---> Translation ---> Protein
Genetic code used during translation:
